How Cell Line Variability Impacts Screening: Cautionary Tale, or Biomarker Boon?
December 28, 2018
Ada Yee
Turning early drug screening hits into successful therapies can be challenging. One possible reason: cell line genetic variability. This article discusses important findings from the Broad Institute of Harvard and MIT showing how variability impacts drug screening, and a possible silver lining.
Variability is the enemy of any biologist struggling in vain to repeat a result. This problem is only amplified in drug discovery, where ensuring the reliability of initial screening results is crucial. An unreliable lead means that huge amounts of time, money and resources could be wasted on developing a drug only to have it fail in clinical trials. What drives variability and what can we do about it?
Cell Lines: Cancer Research Workhorse
To address this question, we’ll look at a recent example from oncology. Human cancer cell lines, basic and long-standing tools of cancer researchers, derive from patient tumor cells. Cell lines can model cellular biology, and because they continually divide, scientists can propagate or passage these cells generation after generation in culture. You’ve probably heard of HeLa, an immortalized cell line derived from a cervical cancer biopsy obtained in 1951 from the patient Henrietta Lacks. Enormously valuable to disease research, cell lines like HeLa have opened the door to discovering many of today’s medicines.
Cell lines are additionally useful for high-throughput drug screening. It would be challenging, expensive or even unethical to perform initial screens of thousands of compounds using in vivo animal models or freshly dissected tissue samples, much less human patients. Cell lines are further amenable to certain types of fluorescent and biochemical assays that wouldn’t be as easily done in these in vivo or ex vivo models. Therefore, cells lines are both a cost-effective and scientifically valuable tool for drug screening.
Another valuable property of cell lines is that every cell generated in the line is a clone. Cell lines are thought to share genetic makeup, making results from a given line comparable from lab to lab. How true is this assumption, however? It is understood that tumor cells become cancerous because they’ve acquired genetic lesions such as single-base pair mutations or loss and duplication of larger blocks of genetic material that enable their abnormal proliferation. Furthermore, with each division, cells copy their DNA, and the copying process is vulnerable to error. This begs the question: How genetically stable are cancer cells lines?
Variability: Enemy to Drug Discovery?
To quantify just how stable cell lines are, Uri Ben-David, Rameen Beroukhim, Todd Golub and colleagues from the Broad Institute of Harvard and MIT recently undertook a heroic task. They performed whole exome sequencing on 106 cancer cells lines grown at the Broad Institute (USA) or at the Sanger Institute (UK).1 Different strains of cell lines have usually been assumed to have uniform genetic makeup, but the researchers found more than the expected genetic variation across different strains of the same cell line, due to mutations and copy number alterations.
The researchers focused on characterizing variation between different strains of a breast cancer cell line called MCF7. They included strains that had been modified with the neutral insertion of fluorescent markers (a typical manipulation that scientists sometimes make to adapt the cell line for a given assay), strains grown in different growth mediums, and one strain that had been previously treated with drug. Ben-David et al. found mutations and copy number variations that affected breast cancer-associated genes such as PTEN and the estrogen receptor gene. Genetic aberrations fell along sub-clonal lines, with more closely-related strains sharing similar alterations. Using live-cell imaging and single-cell RNA sequencing, researchers saw that mutations and other aberrations affected other features of the cells, including size and shape, division rates, and transcriptomic profile (expression of the genes).
What is the consequence of all this variation? The authors performed a screen seeking compounds that inhibited cell growth (as would be desired in a cancer therapeutic). Their screen included 321 compounds which they tested orthogonally across 27 strains; they generated an 8-point dose response curve for each compound × strain permutation, using Genedata Screener® to analyze and fit the data. Of concern, nearly all of the approximate 50 hits showed inconsistent effects from one strain to another, displaying lack of activity in at least one of the strains. This was not due to assay variability, such as variable pipetting or reagents, because replicates and even different compounds with similar known mechanisms of action evoked consistent results between strains. Therefore, the genetic variability within strains had meaningful consequences: by only running your screen in a single cell line, you may risk missing good hits. Worse, you might unknowingly pursue a hit that doesn’t work for many patients. To help scientists better interpret their own experiments, Broad now has created an online tool called Cell STRAINER, which researchers can use to assess how their particular strain diverges genetically from a reference strain, and see how this might alter drug response.
A Silver Lining: Personalized Medicine in Early Discovery?
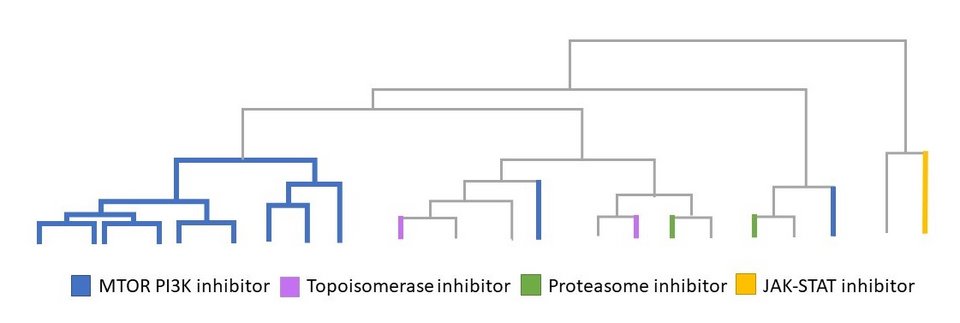
This is not just a cautionary tale, however. The authors observed that the drug response of a particular strain tended to make sense, given the particular genetic quirks of that strain. For example, strains lacking estrogen receptor genes were, as expected, insensitive to estrogen-lowering drugs. To underline this point, the authors used available CRISPR screen data (CEREs), which systemically quantifies the dependency of given cell line strains to given genes.3 Different genetic dependencies indeed correlated with different pharmacological dependencies on given classes of drugs. Thus, the authors proposed that the genetic variability of strains could be harnessed to understand and better target drugs for different patients.
Therefore, a more positive side of cell line variability may be its potential application in biomarker-driven drug development. Granted, it remains unclear how the genetic variations seen in cell lines relates to real patient biomarkers and human genetic variation. Patient-modeling approaches (for example, using patient-derived tissues or stem cells) may still need to complement any cell line screen.2 However, often such materials are non-renewable, and the required protocols more finicky and time consuming than those for cancer cell lines. As a first pass, the genetic variability in cell lines could provide a good proxy for true patient biomarkers.
Increasing Cell Line Screening Throughput
Overall, Ben-David et al.’s work highlights the importance of conducting screens along this second dimension of different cell lines and genotypes. This will require even more high-throughput methods, and analytical software powerful enough to handle this throughput.
Towards this end, the same group at the Broad has developed a technology called PRISM.4 This method enables pooling of several cell lines per plate well by tagging lines with short genetic barcodes for later identification by PCR. Thus, PRISM facilitates and massively increases screening throughput, allowing screening of up to 4,000 compounds across 600 cells lines, generating over 700,000 dose-response curves. Genedata has demonstrated that Genedata Screener can handle such large-scale PRISM data, automating quality control and allowing you to generate dose-response curves and other analyses (Request poster, "Large-scale testing of compounds across the diversity of human cancer types could become a routine activity".)
Conclusion
While this case concerns cancer research, cell lines are used to study many diseases, so these lessons may transfer to other therapeutic areas. To guard against misinterpretation of cell line data, screening across multiple, carefully characterized cell line strains will become a necessary part of drug discovery. Furthermore, screening across cell line strains could even be a boon to personalized medicine, bringing biomarker-based approaches earlier into the screening process.
References:
1. Ben-David et al. " Genetic and transcriptional evolution alters cancer cell line drug response." Nature 560: 325–330 (2018).
2. Shi et al. "Induced pluripotent stem cell technology: a decade of progress." Nature Reviews Drug Discovery 16:115-130 (2017).
3. Meyers et al. "Computational correction of copy-number effect improves specificity of CRISPR-Cas9 essentiality screens in cancer cells." Nature Genetics 49:1779-1784 (2017).
4. Yu et al. "“High-throughput identification of genotype-specific cancer vulnerabilities in mixtures of barcoded tumor cell lines." Nature Biotechnology 34:419-423 (2016).
Author: Ada Yee, Ph.D., Scientific Story Researcher, Genedata Screener.