論文掲載:ディープラーニングを使用したHCS画像解析
May 22, 2020
Stephan Steigele
I’m pleased to share with you that our paper is out, in the SLAS Discovery SBI2 special issue. The paper talks about Genedata Imagence®, novel AI-based enterprise software, and describes its real-world approach for using AI in high-content screening (HCS) image analysis.
While you can already learn quite a bit about Genedata Imagence at Genedata’s website, I also want to give my personal take on this solution and share our experiences building and testing it with customers.
Our Journey to Building Genedata Imagence
At the time we first came up with the idea behind Genedata Imagence almost three years ago, we already had a lot of expertise in high-content image data analysis: our platform for HCS analysis, Genedata Screener® for HCS, had been available to our customers for many years. At that point, we had been thinking for some time about how to enhance this HCS offering. We experimented quite a lot with state-of-the-art machine learning, with the goal of making HCS even more efficient and productive for our users. This search for a much more efficient high-content analysis workflow was also motivated by a recent sharp increase in phenotypic screening—especially with the rise of new assay formats like the Cell Painting assay and the anticipation that these would be adopted quite quickly by the industry. With the rapid advancement of AI algorithms and tools, and the particular success of AI in image recognition, it became clear we had a golden opportunity to combine our expertise in drug discovery and data analytics to create a novel solution for HCS scientists.
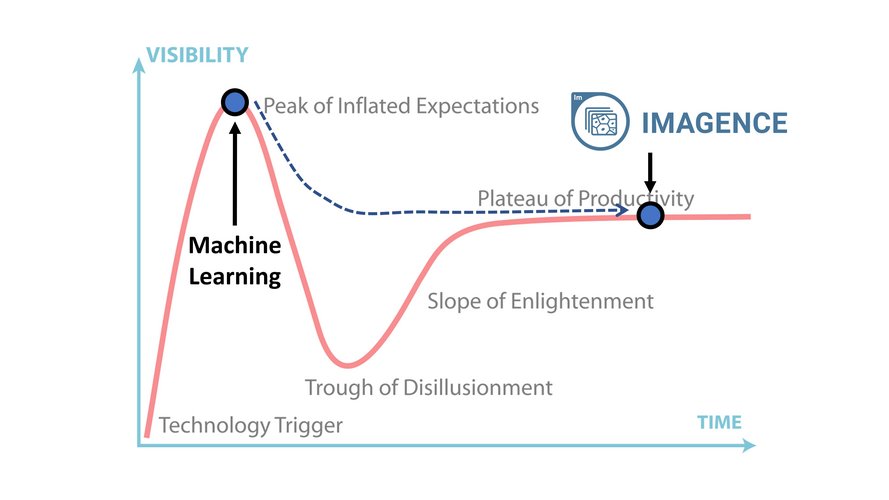
We also realize there is a lot of hype surrounding AI and its use in drug discovery. Gartner’s 2019 Hype Cycle puts machine learning at the apex of the Peak of Inflated Expectations, but behind this hype is the promise that this new technology will yield great success. Our goal was to create a solution that advances the use of ML and AI in drug discovery all the way to the Plateau of Productivity.
We knew from the get-go that we wanted our solution to be (1) an out-of-the box solution that eliminates any barriers to using powerful deep learning tools for biologists without training in AI, and (2) a practical solution that integrates into an enterprise, production setting.
The former we accomplished in part by using “Similarity Map” visualizations that enable intuitive result exploration, and in part through highly automated phenotype classification and feature detection afforded by a tailored deep learning approach. The latter we have accomplished using highly parallelized processing, automated image upload, and facilitated reporting to data warehouses. Both these aspects of our solution are showcased and explained in our paper.
In hands-on sessions with multiple customers who took part in the Early Access program for Genedata Imagence , I’ve had the great satisfaction of seeing firsthand how HCS biologists could really dig into the Similarity Maps without the need for any extensive training, letting their own biological knowledge guide them. We’ve been both impressed and intrigued to see the wide range of assays to which users applied our software, including everything from toxicity studies to Cell Painting assays and MoA analyses. And of course, we were honored when our product won the Best in Show Bio-IT Award in 2019 (in the “No BS AI” category), for the real-world applicability of our solution.
New Frontiers: Transferring Insight Using Genedata Imagence
Also addressed in the paper is how Imagence allows transfer of knowledge gained in one assay to distinct yet similar assays. This feature was originally born out of a collaboration with Astra Zeneca that also later won a BioIT Best in Show Award, in 2018. Genedata Imagence provides a simple workflow that makes it easy for those not versed in AI to employ this more advanced technique to their HCS datasets, again using the concept of expert visual guidance via Similarity Maps. We think this will be a particularly powerful tool for scientists, allowing them to apply prior knowledge in an unbiased way and freeing up their time for new tasks instead of assay re-development.
A New Model for AI in Drug Discovery
Another important aspect of our paper is that it outlines a model for the way AI can be applied in drug discovery more broadly. While HCS provided a particularly good starting point for building an enterprise-ready AI-based discovery solution, we also think that Genedata Imagence can serve as a blueprint for how AI can be most productively applied at scale in enterprise settings. This includes the use of visualization to aid with insight, automation, multi-level quality control, and integration with enterprise software. The team at Genedata is actively working on new ideas for how to integrate AI into discovery workflows, and I’m excited to see these projects bear fruit.
Speaking of teams—I want to end this post by thanking my colleagues Daniel Siegismund, Matthias Fassler, and Marusa Kustec for their pivotal work in the initial development of Genedata Imagence and their continued drive to evolve it further and share it with new users. I also thank the Genedata software development and management teams (especially, Annette Brodte, Bernd Kappler, and Stephan Heyse) for their crucial role in making Genedata Imagence a finished product. I can’t wait to see what future users of Genedata Imagence will be able to accomplish with it!
Stephan Steigele, Ph.D., is Head of Science, Genedata Screener